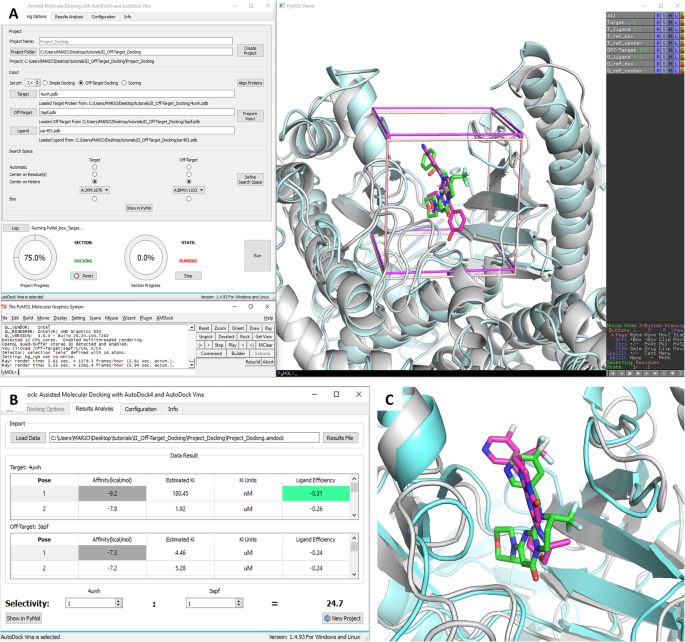
AMDock: a versatile graphical tool for assisting molecular docking with Autodock Vina and Autodock4 | Biology Direct | Full Text

AutoDock Vina 1.2.0: New Docking Methods, Expanded Force Field, and Python Bindings | Journal of Chemical Information and Modeling

Molecular Docking | Autodock VINA Virtual Screening | VINA Docking tutorial | Bioinformatics - YouTube