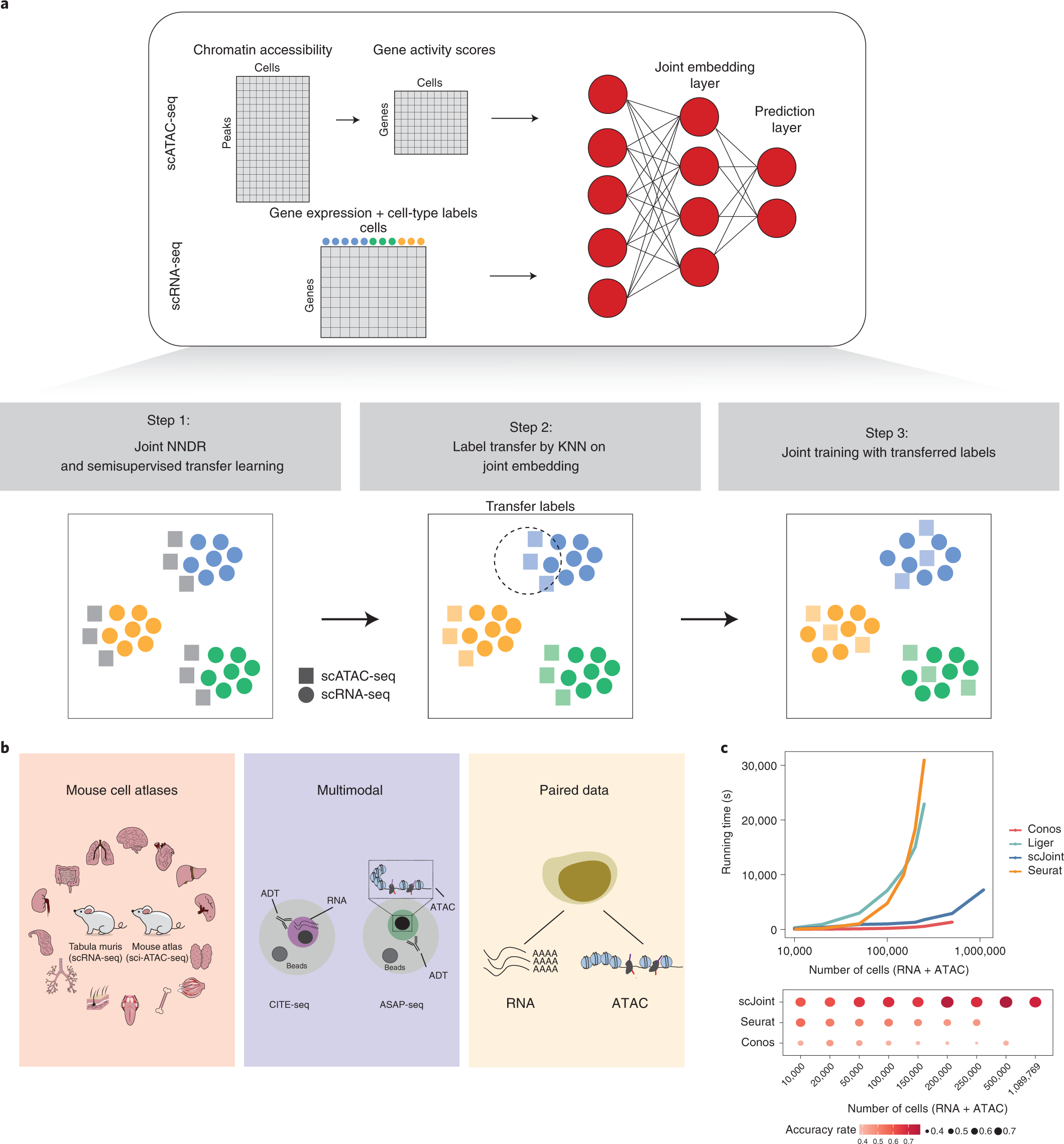
scJoint integrates atlas-scale single-cell RNA-seq and ATAC-seq data with transfer learning | Nature Biotechnology
Milena Hasan - Jost Enninga - Single Cell Resources Initiative at Institut Pasteur - Research - Institut Pasteur

Whole-animal multiplexed single-cell RNA-seq reveals transcriptional shifts across Clytia medusa cell types | Science Advances
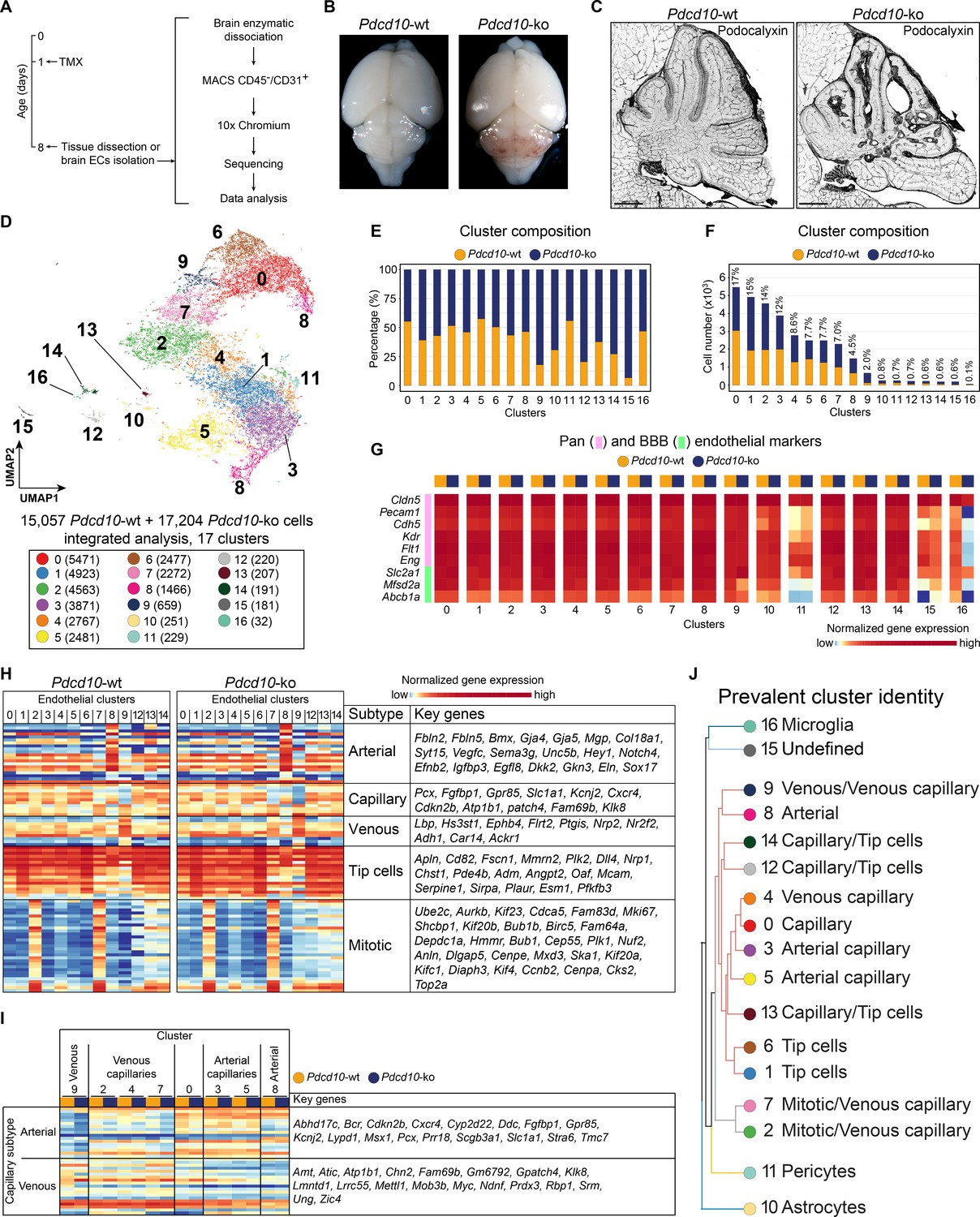
Mapping endothelial-cell diversity in cerebral cavernous malformations at single-cell resolution | eLife
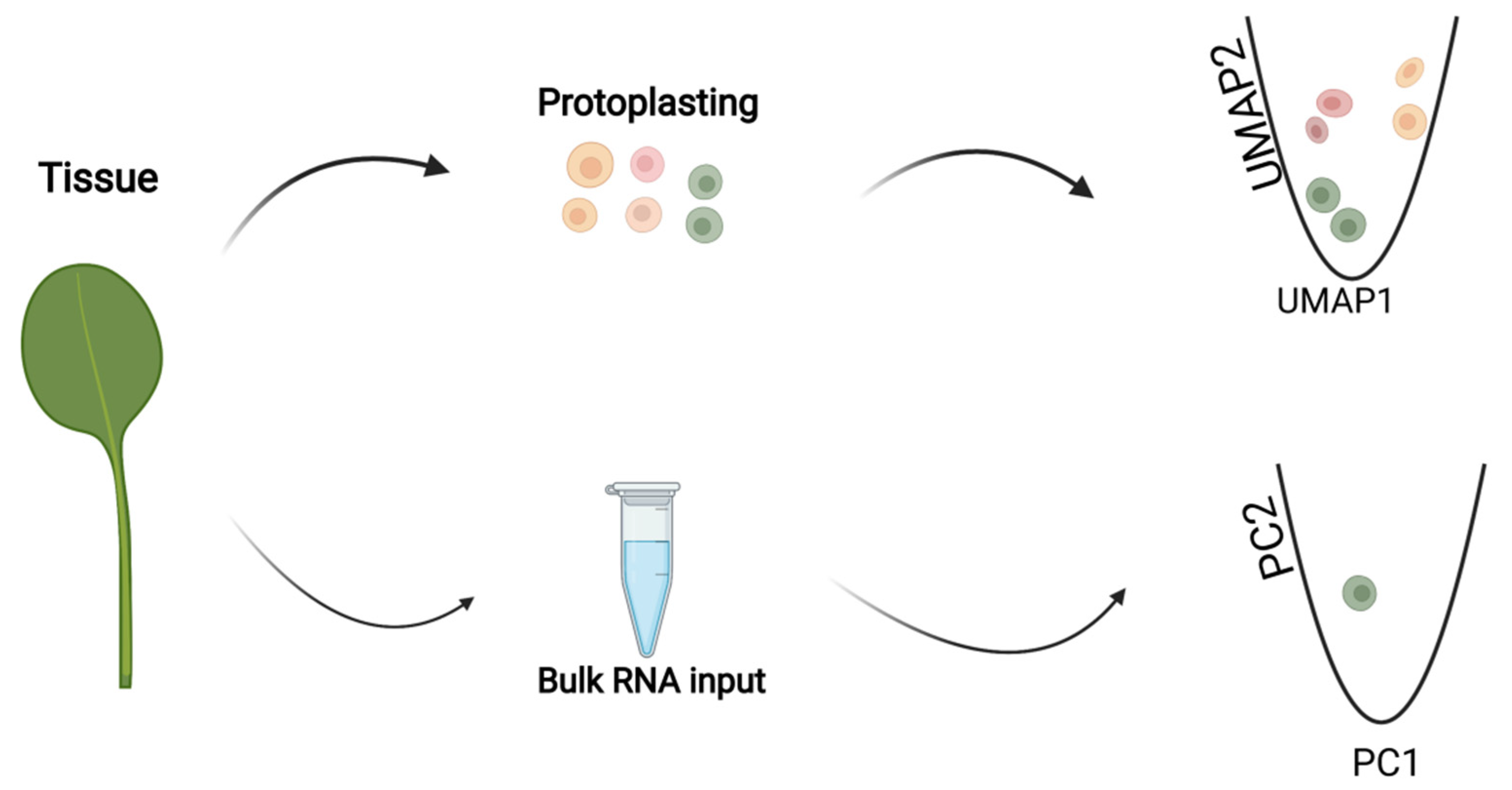
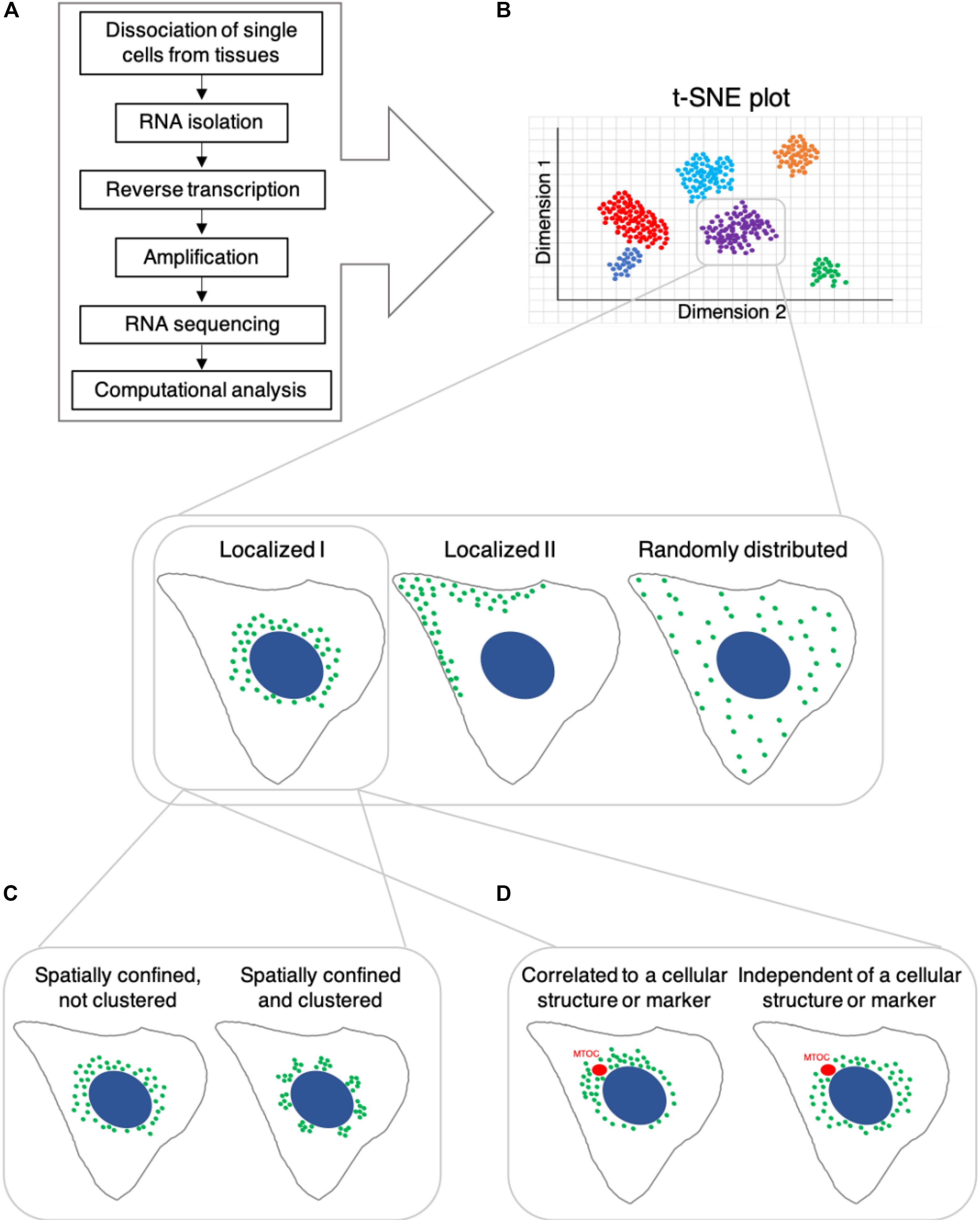
IJMS | Free Full-Text | Single-Cell RNA Sequencing for Plant Research: Insights and Possible Benefits

The Repertoire of Serous Ovarian Cancer Non-genetic Heterogeneity Revealed by Single-Cell Sequencing of Normal Fallopian Tube Epithelial Cells: Cancer Cell
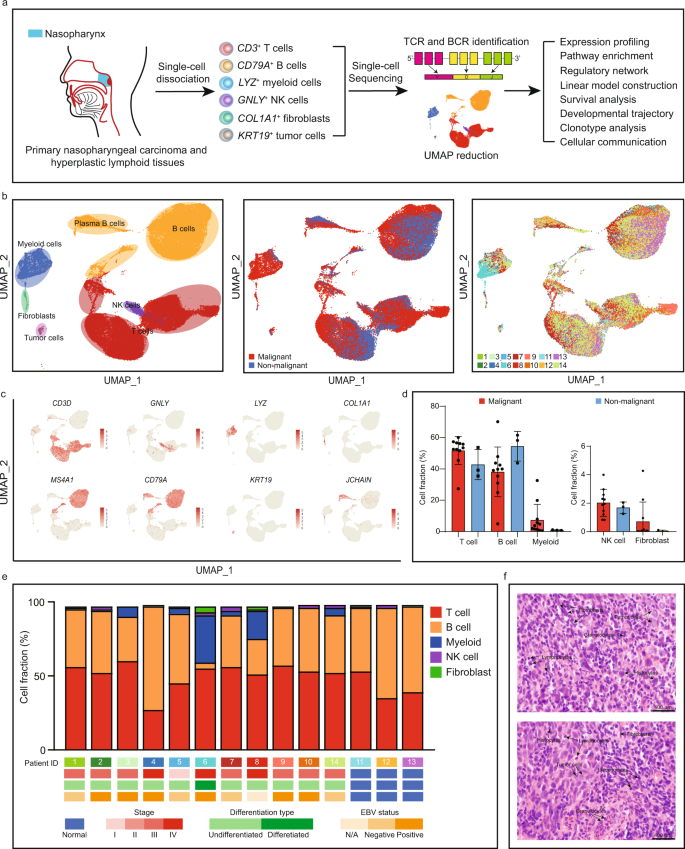
Comprehensive single-cell sequencing reveals the stromal dynamics and tumor-specific characteristics in the microenvironment of nasopharyngeal carcinoma | Nature Communications

PDF) Comprehensive network modeling from single cell RNA sequencing of human and mouse reveals well conserved transcription regulation of hematopoiesis
Scedar: A scalable Python package for single-cell RNA-seq exploratory data analysis | PLOS Computational Biology
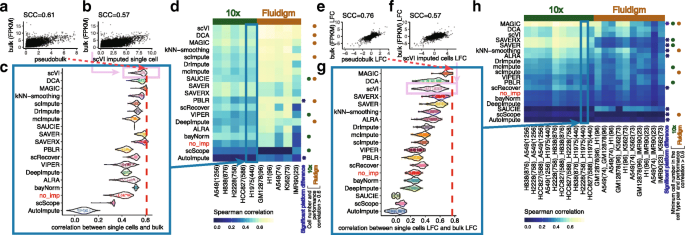
A systematic evaluation of single-cell RNA-sequencing imputation methods | Genome Biology | Full Text
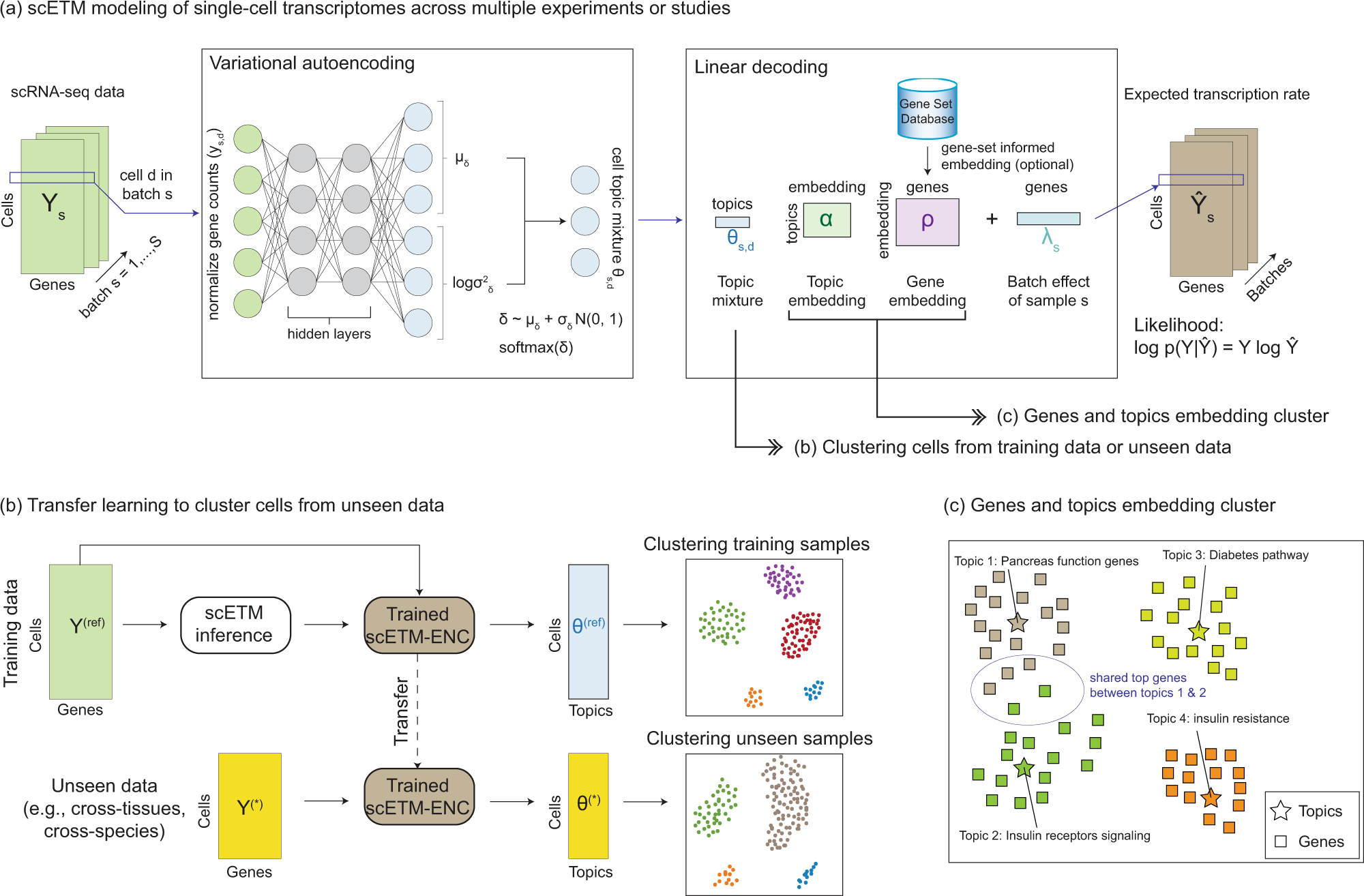
Learning interpretable cellular and gene signature embeddings from single- cell transcriptomic data | Nature Communications

Benchmarking Computational Doublet-Detection Methods for Single-Cell RNA Sequencing Data - ScienceDirect
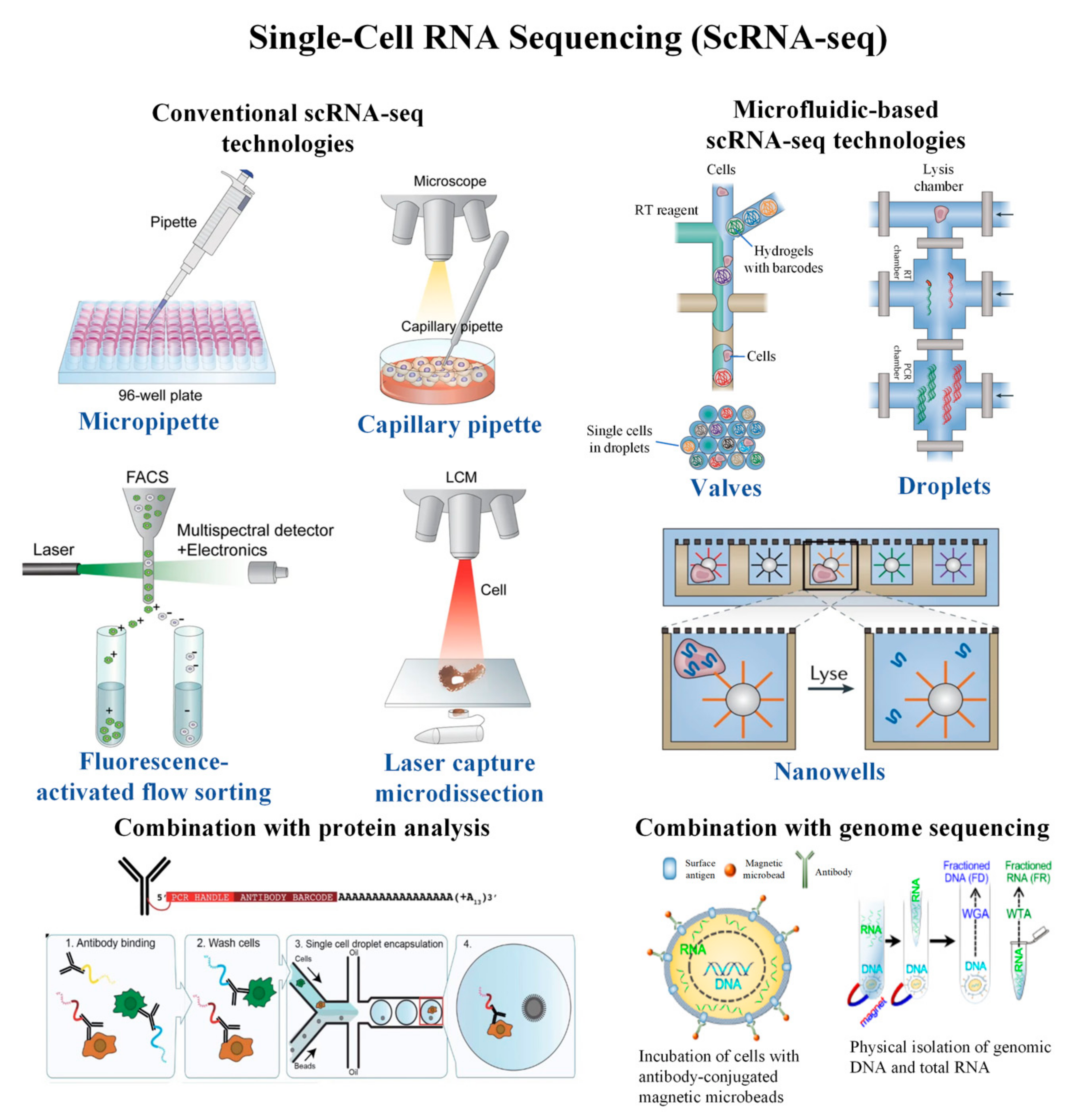
Cells | Free Full-Text | Single-Cell RNA Sequencing and Its Combination with Protein and DNA Analyses
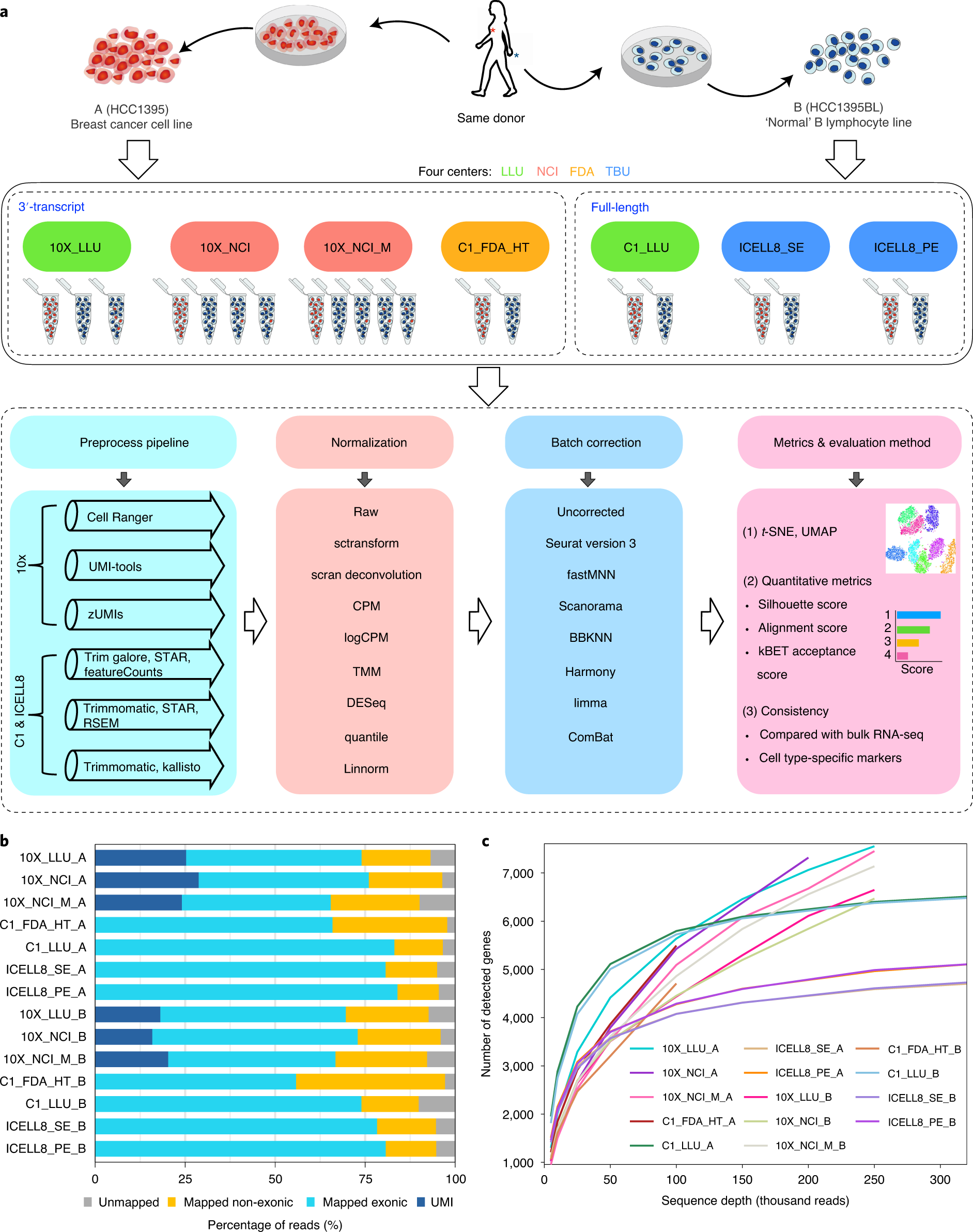
A multicenter study benchmarking single-cell RNA sequencing technologies using reference samples | Nature Biotechnology

Single Cell Sequencing Reveals Glial Specific Responses to Tissue Processing & Enzymatic Dissociation in Mice and Humans | bioRxiv

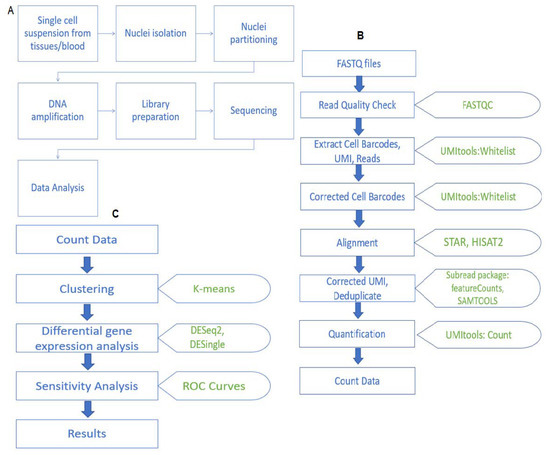
Single-cell ATAC sequencing analysis: From data preprocessing to hypothesis generation - ScienceDirect
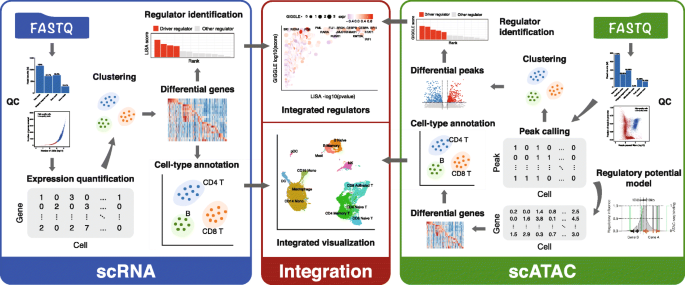
Integrative analyses of single-cell transcriptome and regulome using MAESTRO | Genome Biology | Full Text